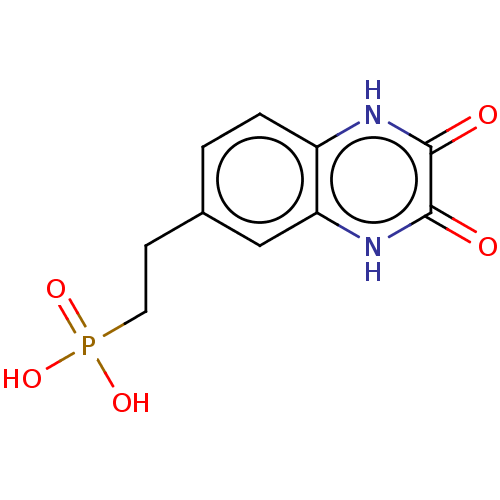
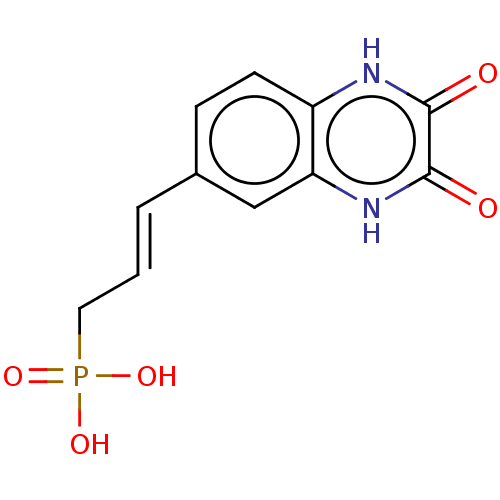
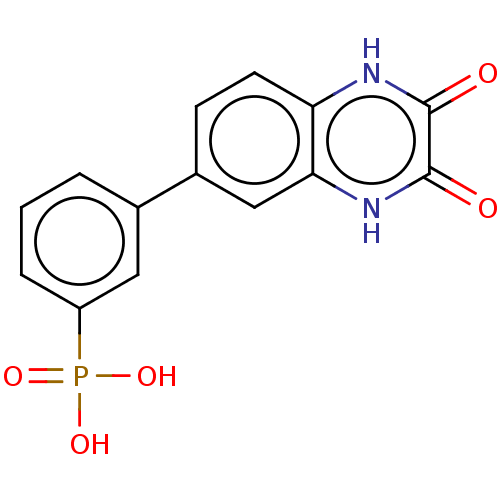
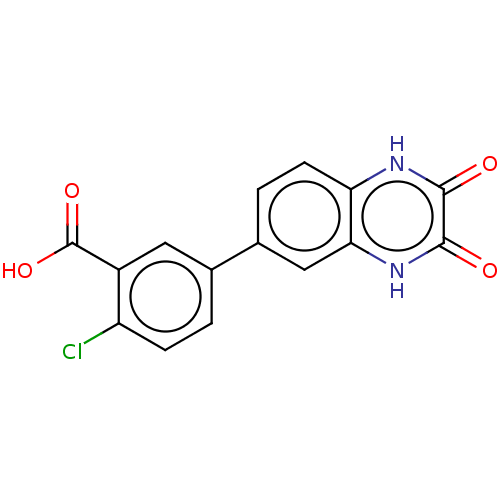
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
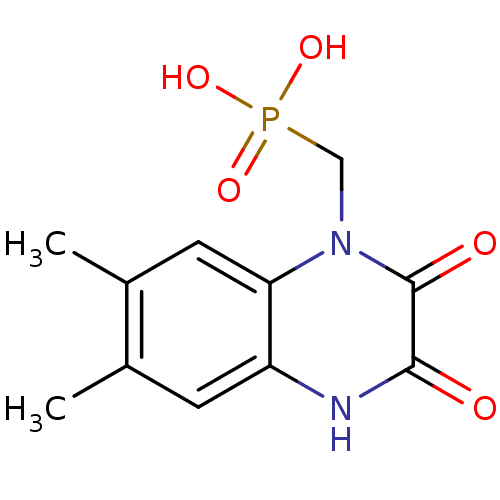
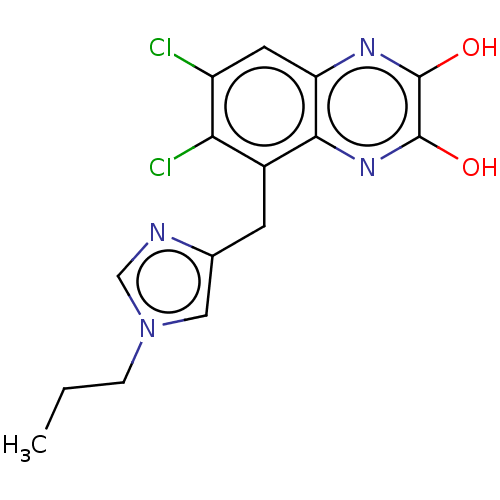
Affinity DataKi: 0.960nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
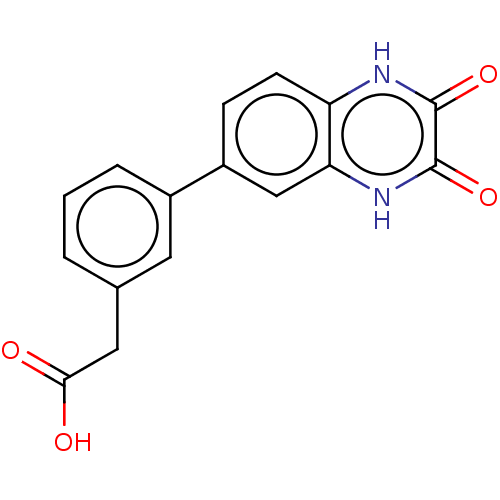
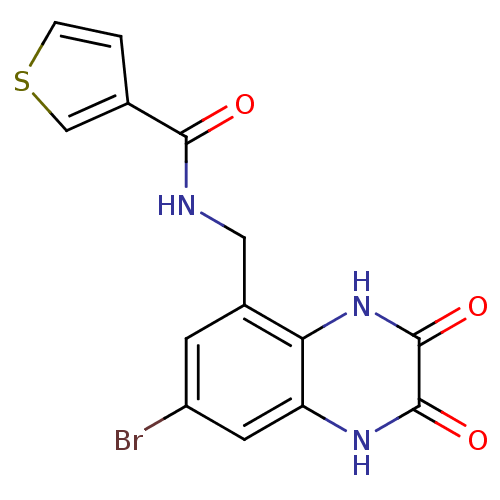
Affinity DataKi: 1.40nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
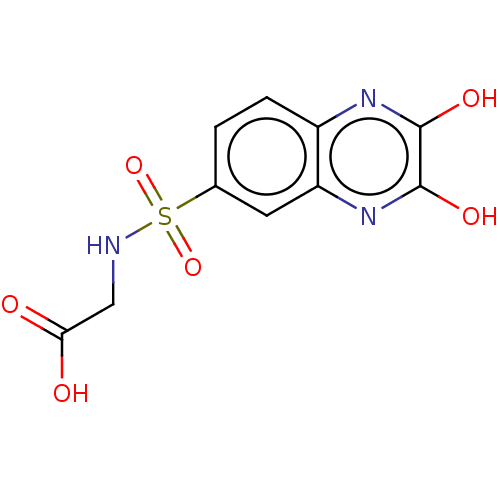
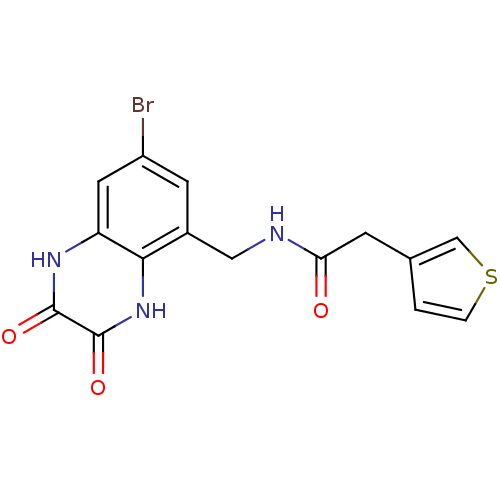
Affinity DataKi: 2.30nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
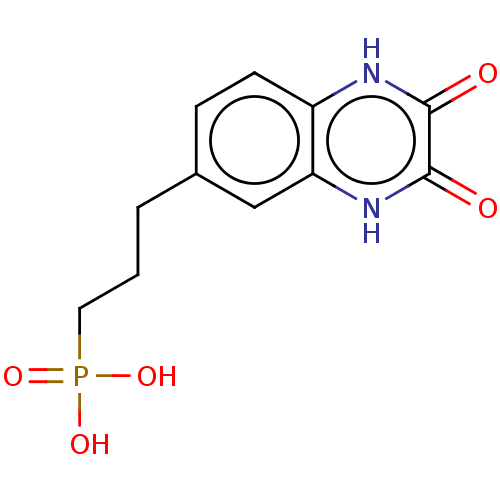
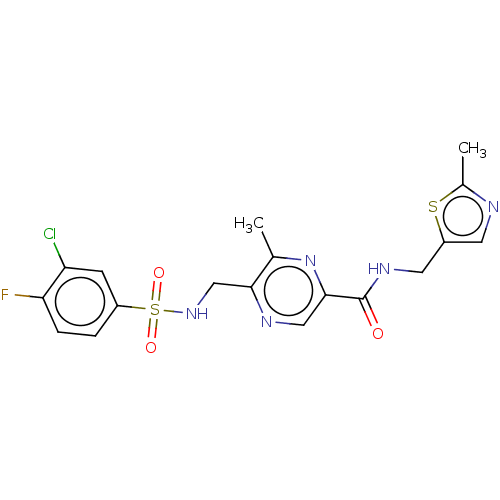
Affinity DataKi: 2.60nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 3.10nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 3.30nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 4.40nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 5.10nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 6.60nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 9.60nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 9.90nMAssay Description:Displacement of [3H]DCKA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 65nM ΔG°: -41.0kJ/mole IC50: 92nMpH: 7.4 T: 2°CAssay Description:Two-electrode voltage clamp (TEVC) recordings were performed on Xenopus oocytes at room temperature 3-6 days postinjection using an OC-725C TEVC ampl...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Displacement of [3H]glycine from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Ligand Info
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Ligand Info
Affinity DataKi: 560nM ΔG°: -35.7kJ/mole IC50: 770nMpH: 7.4 T: 2°CAssay Description:Two-electrode voltage clamp (TEVC) recordings were performed on Xenopus oocytes at room temperature 3-6 days postinjection using an OC-725C TEVC ampl...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 590nMAssay Description:Binding affinity to NMDAR (unknown origin) assessed as inhibition constant by [3H]MK-801 binding assayMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 1.90E+3nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 5.28E+3nMAssay Description:Displacement of [3H]glycine from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 1.38E+4nMAssay Description:Displacement of [3H]glycine from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 4.10E+4nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 5.30E+4nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 5.50E+4nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 5.60E+4nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: 6.90E+4nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 2.60nMAssay Description:Displacement of [3H]L-689560 from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 4.40nMAssay Description:Displacement of [3H]L-689560 from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Displacement of [3H]MDL from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Displacement of [3H]MDL from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Displacement of [3H]MDL from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 16.2nMAssay Description:Cell Culture and plating: HEK293 cells expressing NR1/NR2A (Chantest, Cleveland, Ohio) were grown to 70-80% confluency as an adherent monolayer in st...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 19.6nMAssay Description:Cell Culture and plating: HEK293 cells expressing NR1/NR2A (Chantest, Cleveland, Ohio) were grown to 70-80% confluency as an adherent monolayer in st...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Antagonist activity at NR2A transfected in oocytesMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 35nMAssay Description:Inhibition of [3H]CPP binding to rat N-methyl-D-aspartic acid receptor 2AMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 39.5nMAssay Description:Cell Culture and plating: HEK293 cells expressing NR1/NR2A (Chantest, Cleveland, Ohio) were grown to 70-80% confluency as an adherent monolayer in st...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 61nMAssay Description:Cell Culture and plating: HEK293 cells expressing NR1/NR2A (Chantest, Cleveland, Ohio) were grown to 70-80% confluency as an adherent monolayer in st...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 70nMAssay Description:Binding affinity to NMDA receptor (unknown origin)More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 87.8nMAssay Description:Cell Culture and plating: HEK293 cells expressing NR1/NR2A (Chantest, Cleveland, Ohio) were grown to 70-80% confluency as an adherent monolayer in st...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 90nMAssay Description:Displacement of [3H]NMDA from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 95nMAssay Description:Inhibition of [3H]CPP binding to rat N-methyl-D-aspartic acid receptor 2AMore data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Inhibitory activity against N-methyl-D-aspartate glutamate receptor 2A in ratMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2A(Homo sapiens (Human))
Central South University
Curated by ChEMBL
Central South University
Curated by ChEMBL
Affinity DataIC50: 108nMAssay Description:Cell Culture and plating: HEK293 cells expressing NR1/NR2A (Chantest, Cleveland, Ohio) were grown to 70-80% confluency as an adherent monolayer in st...More data for this Ligand-Target Pair









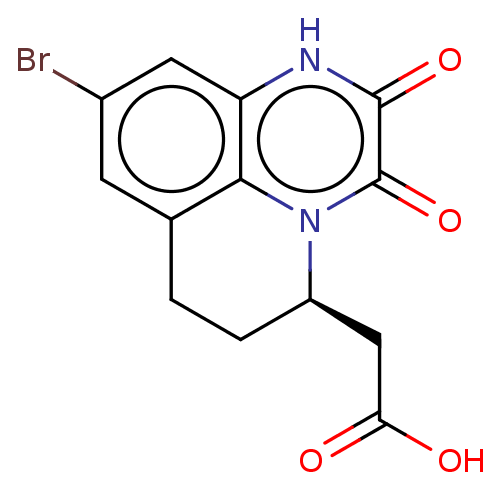

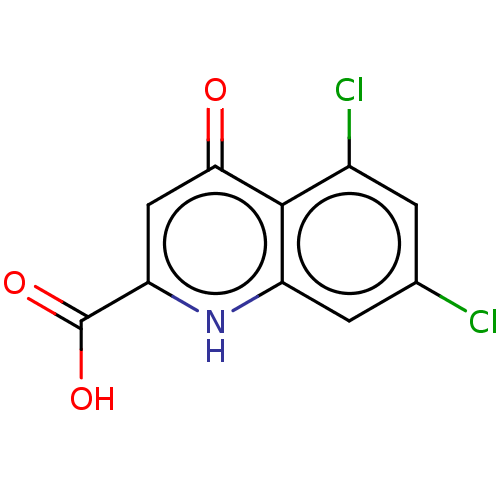
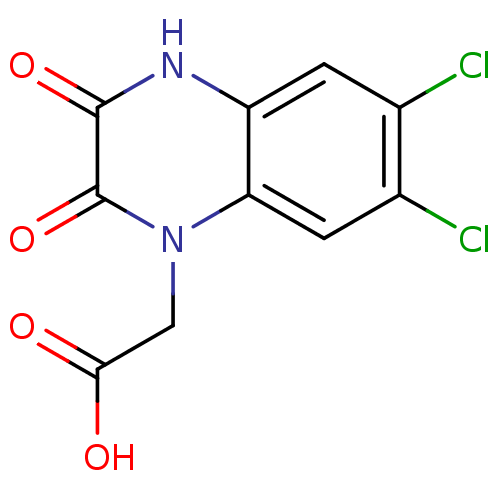
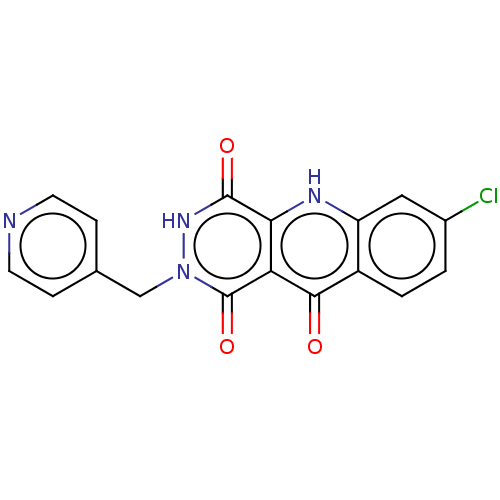




 3D Structure (crystal)
3D Structure (crystal)